
SIMULTANEOUS CFDNA AND EXOSOME ISOLATION FROM THE SAME SAMPLE (PLASMA, URINE)
SubXTM-Exo-DNA kit is designed for simultaneous isolation of exosomes and circulating cell-free DNA (cfDNA) directly from liquid biopsies (plasma or serum) without Proteinase K. Our proprietary bi-functional substance (SubXTM) binds DNA under physiological conditions (e.g. directly in biological liquids) with high affinity, followed by adsorption of the [DNA- SubXTM] complex on a solid phase matrix. SubXTM captures DNA via its phosphate groups and eliminates any bias related to both AT/GC content and DNA fragments length improving extraction efficacy and accuracy of downstream applications. Since our method does not require chaotropic salts for DNA capture from the starting material, no dilution of exosomes and cfDNA takes place. As both ends of the SubXTM molecule have phospholipid binding groups, each molecule of SubXTM can anchor two exosomes and thus produce dimers and then oligomers of the extracellular vesicles. SubXTM can oligomerize exosomes in both highly proteinaceous bioliquids (serum, plasma) and in high salt biofluids (urine, tears, etc.). Excess of SubXTM results in oligomerization of up to 10-15 exosomes and formation of micron-size particles that are easily precipitated in a brief 14K x g centrifugation step. A specially designed buffer allows for reconstitution of the pelleted exosomes back to free monomer state suitable for downstream applications. Different affinity of SubXTM to DNA (strong) and to exosome phospholipid groups (mild) allows for separation of the exosomes and cfDNA from the total pellet. Since DNA molecule has a high density of phosphate residues, [cfDNA+SubX] complex is very stable. In contrast, [exosomes+SubX] complex posses a middle stability due to lower density of phosphate residues on the vesicles outer membrane. Exosomes are easily extracted from the pellet by low salt reconstitution buffer while cfDNA remains tightly bound with SubX and remains insoluble. Specially designed high salt buffer is required to deliberate cfDNA from the SubX complex/pellet. As a result, free circulating cfDNA does not contaminate exosomal preparations. Thus, two important subsets of extracellular molecules/microvesicles – cfDNA and exosomes - can be separated in a single protocol without cross-contamination.
The following diagram shows the sequence of steps involved in the process of separation and isolation of pure exosomes and cfDNA even from the small volume of the starting material (plasma or serum).
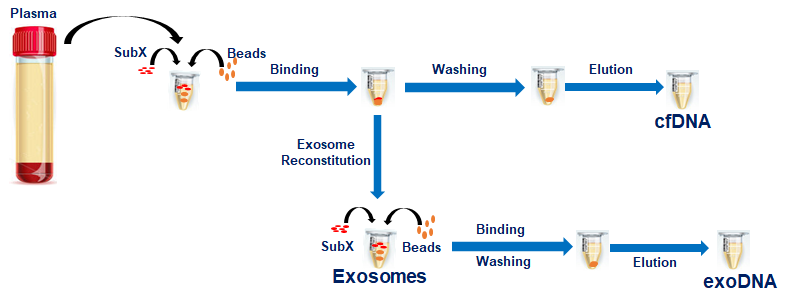
Figure 1. Exosome and DNA isolation steps. All procedures after mixing plasma or urine with SubX are done in one tube. Each incubation takes place at room temperature for 3-4 minutes with occasional vortexing. Both centrifugation or magnet separation can be used to remove solutions. Exosomes are separated from SubX-DNA-Beads pellet by reconstitution in a special buffer ERB. DNA is eluted from the beads at 60°C.
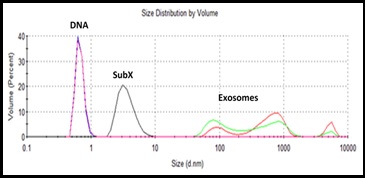
Figure 2. Simultaneous binding of urine endogenous exosomes and sonicated spike DNA with SubXTM followed by sequential isolation of exosomes and then DNA. Profiles of spike DNA (blue) and exosomes (green) before binding with SubXTM and after dissociation (pink and red) perfectly coincide.
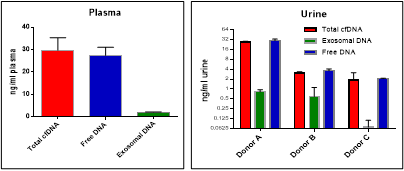
Figure 3. cfDNA isolated from normal donor plasma and urine. Over 90% cfDNA persists in a free circulating form and only traces of cfDNA are associated with exosomes. DNA yield was measured by Qubit HS DNA Assay.

